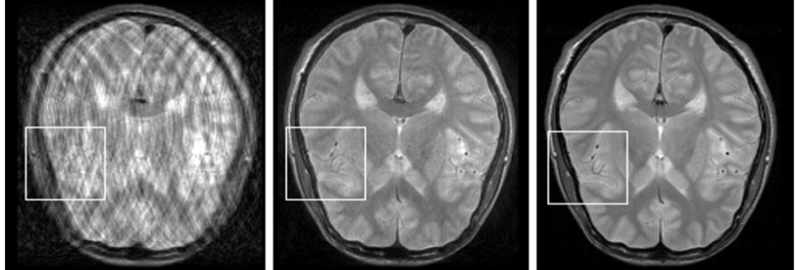
Retrospective, fully data-driven removal of motion-related artefacts in 2D images acquired with Turbo Spin-Echo sequence. Left: motion corrupted image. Middle: reconstruction. Right: no-motion image.
Modern medical imaging is a blend of 20th-century breakthroughs in physics and information processing. Although there is an ongoing advancement of technology, many of the problems are still unsolved. Using machine learning and image processing techniques for more efficient and intelligent use of acquired data seems a promising solution to some of the obstacles. For example, understanding the statistical properties of medical images allows making certain prior assumptions on unobserved or corrupted aspects of the data, which makes it possible to improve the image quality and aid the medical diagnostics.
PET-MR systems combine functional information from Positron Emission Tomography (PET) with structural information from Magnetic Resonance (MR) Imaging. They require a PET attenuation (or $\mu$) map to be determined from MR data, which constitutes an ill-posed problem. We have developed a Gaussian process prediction framework, trained on co-registered MR and $\mu$-maps, and evaluated it on brain [ ] and whole body [ ] data. The research paper which first described the method [ ] received a Best Basic Science Investigation Award of the Journal of Nuclear Medicine in 2008. A collaborative effort with the university hospital, Tübingen, the method has been patented and licensed to Siemens AG. Further research focused on evaluating the influence of positioning aids [ ], improving the robustness of the method in presence of MR image artifacts [ ] and the assessment of PET quantification accuracy in a pediatric patient collective [ ].
Patient motion during long MR scans leads to severe non-local degradations and can render images unacceptable for medical diagnosis. We developed a retrospective (post-processing) approach called GradMC, which operates on the motion-corrupted images [ ]. It automatically corrects for motion artifacts by searching for a motion-generating function whose inverse application optimizes a quality measure based on image statistics. Our GPU implementation processes a full 3D volume in a few minutes, which is acceptable in routine clinical use. In follow-up work, we have come up with a fully retrospective non-rigid motion correction scheme that only needs raw data as an input [ ].
Diffusion MRI (dMRI) allows us to investigate brain tissue microstructure non-invasively. Recently, supervised classification has been used to recognize patterns in this rich and complex data that allow us to automatically detect neurological diseases. We have contributed to this field by developing improved methods for feature extraction [ ], and by applying machine learning to specific diseases [ ].
